Introduction¶
Table of contents
pychemcurv is a python package for structural analyzes of molecular systems or solid state materials focusing on the local curvature at an atomic scale. The local curvature is then used to compute the hybridization of molecular orbitals.
The main features of the library are available from a Plotly/Dash web application available here: pychemcurv.onrender.com/. The web-app allows to upload simple xyz files and compute the local geometrical properties and the hybridization properties. The application source code is available in a separate repository at pychemcurv-app.
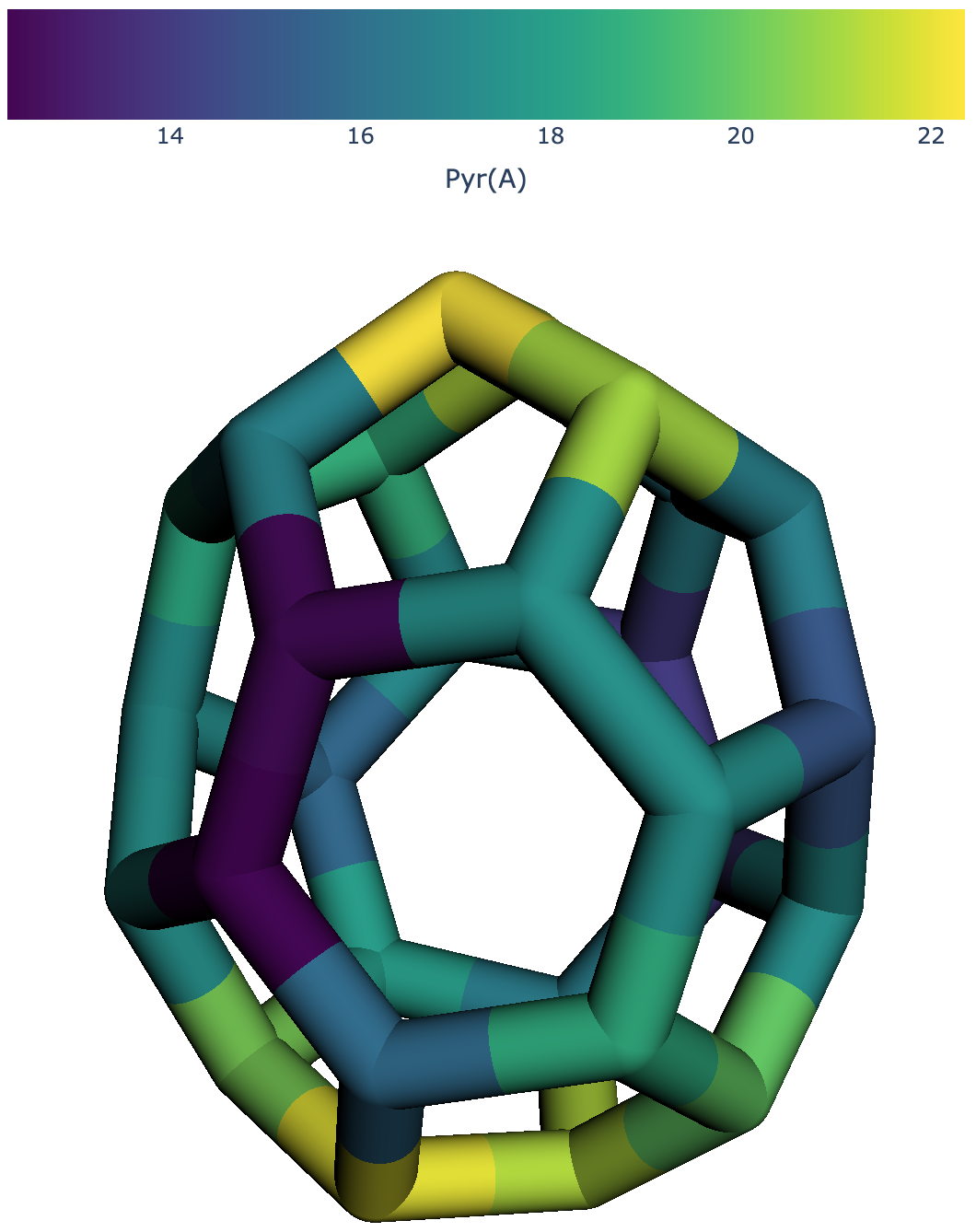
Pyramidalization angle of a \(C_{28}\) fullerene mapped on the structure with a colorscale.¶
Features¶
Pychemcurv is divided in two parts. The first one is a standard python package which provides two main classes to compute the local curvature at the atomic scale and the hybridization of a given atom. Second, a Plotly/Dash web application is provided in order to perform a geometrical and electronic analyzes on molecules or materials.
The web application is available at pychemcurv.onrender.com/. The web-app allows to upload simple xyz files and compute the local geometrical properties and the hybridization properties. The application source code is available in a separate repository at pychemcurv-app.
Some jupyter notebooks are provided in the notebooks/ folder and present use cases
of the classes implemented in this package. You can access to these notebooks
online with binder.
Citing pychemcurv¶
Please, consider to cite the following papers when using either the pychemcurv library or the web application.

Julia Sabalot-Cuzzubbo, Germain Salvato Vallverdu, Didier Bégué and Jacky Cresson Relating the molecular topology and local geometry: Haddon’s pyramidalization angle and the Gaussian curvature, J. Chem. Phys. 152, 244310 (2020).

Julia Sabalot-Cuzzubbo, N. Cresson, Germain Salvato Vallverdu, Didier Bégué and Jacky Cresson Haddon’s POAV2 vs POAV theory for non-planar molecules, J. Chem. Phys. 159, 174109 (2023).
Installation¶
Installation from PyPi¶
From November 2023, pychemcurv is available on pypi. You can install it
directly using pip.
python -m pip install pychemcurv
Installation from source¶
Before installing pychemcurv it is recommanded to create a virtual environment
using conda or virtuelenv.
In this environment, using pip directly from the github repository, run
pip install git+git://github.com/gVallverdu/pychemcurv.git
Alternatively, you can first clone the pychemcurv repository
git clone https://github.com/gVallverdu/pychemcurv.git
and then install the module and its dependencies using
pip install .
If you want to use the web application locally or if you want to use
nglview to display structures in
jupyter notebooks you need to install more dependencies. The setup configuration
provides the viz extra so, using pip, run one of
pip install .[viz]
# escape square bracket with zsh
pip install .\[viz\]
If you have installed nglview you may have to enable the jupyter extension
jupyter-nbextension enable nglview --py --sys-prefix
Install in developper mode¶
In order to install in developper mode, first create an environment (using one of the provided file for example) and then install using pip
pip install -e .[viz]
If you want to build the documentation you also need to install sphinx.
A dedicated requirements file is provided in the docs/ folder.
Run the web application¶
The web application is available in this separate repository: pychemcurv-app https://github.com/gVallverdu/pychemcurv-app. The main aim of the application is to use the pychemcurv package and visualize the geometrical or chemical atomic quantities mapped on the chemical structure of your system.
The application is available online at this address: pychemcurv.onrender.com/.
Demo video:
In order to run the application locally, you have to clone the repository and
install all the dependencies. In particular dash and dash-bio.
You can do that from the requirements.txt provided in the repository of the
application. Here is a short procedure in order to install and run the application
locally. It assumes pychemcurv is already installed in a python environment
called (curv):
[user@computer] (curv) > $ git clone https://github.com/gVallverdu/pychemcurv-app.git
[user@computer] (curv) > $ cd pychemcurv-app/
[user@computer] (curv) > $ python -m pip install -r requirements.txt
[user@computer] (curv) > $ python app.py
Running on http://127.0.0.1:8050/
Debugger PIN: 065-022-191
* Serving Flask app "app" (lazy loading)
* Environment: production
WARNING: This is a development server. Do not use it in a production deployment.
Use a production WSGI server instead.
* Debug mode: on
Open the provided url to use the application.
You can switch off/on the debug mode by setting debug=False on the last line of
the app.py file.
Common error on local execution¶
If the application does not start with an error such as:
socket.gaierror: [Errno 8] nodename nor servname provided, or not known
Go to the last lines of the file app.py and comment/uncomment the last lines to get something that reads
if __name__ == '__main__':
app.run_server(debug=True, host='127.0.0.1')
# app.run_server(debug=False)
Licence and contact¶
This software was developped at the Université de Pau et des Pays de l’Adour (UPPA) in the Institut des Sciences Analytiques et de Physico-Chimie pour l’Environement et les Matériaux (IPREM) and the Institut Pluridisciplinaire de Recherches Appliquées (IPRA) and is distributed under the MIT licence.
Authors
Germain Salvato Vallverdu: germain.vallverdu@univ-pau.fr
Julia Sabalot-cuzzubbo julia.sabalot@univ-pau.fr
Didier Bégué: didier.begue@univ-pau.fr
Jacky Cresson: jacky.cresson@univ-pau.fr

